About Me

Hello! I'm Natalie Davidson, a Post-Doctoral researcher in the Greene Lab at the University of Colorado School of Medicine. I received my Ph.D. from Tri-Institutional Program for Computational Biology and Medicine in New York, while conducting research at ETH Zürich.
I am interested in building methods to integrate multi-scale and diverse data sources to better understand disease. For example, while single-cell experiments provide greater resolution within a single sample, they lack coverage across diseases, tissues, and perturbations. In contrast, bulk data loses cell-specific information, but currently covers more diverse sets of donors. Using both data types can help us make the most use of the available datasets, which is especially beneficial for rare diseases. The methods I am most excited to explore in this space are structurally restricted neural networks. Through careful design of these networks, we can leverage our existing biological knowledge of cells, tissues, signaling pathways, and experimental design to obtain more generalizable and interpretable models.
Prior Work
During my Ph.D., I conducted single cancer and pan-cancer analyses using RNA-Seq and Ribosome Profiling data. I typically use generalized linear models and generalized linear mixed models. I was a part of the International Cancer Genome Consortium, where I integrated multiple transcriptional aberrations such as splicing, fusions, over/under expression, allele specific expression, and others in over 1,000 samples to identify cancer relevant genes and alteration patterns. I received my B.Sc. in computer science and minor in applied mathematics from University of California, Santa Barbara. I received my M.Sc. in computer science from University of California, Los Angeles under the advisement of Dr. Jason Ernst.
Thesis Title: "Identification and Characterization of Aberrations in Cancer through the lens of RNA." (link)

Code
Presentations
(full publications)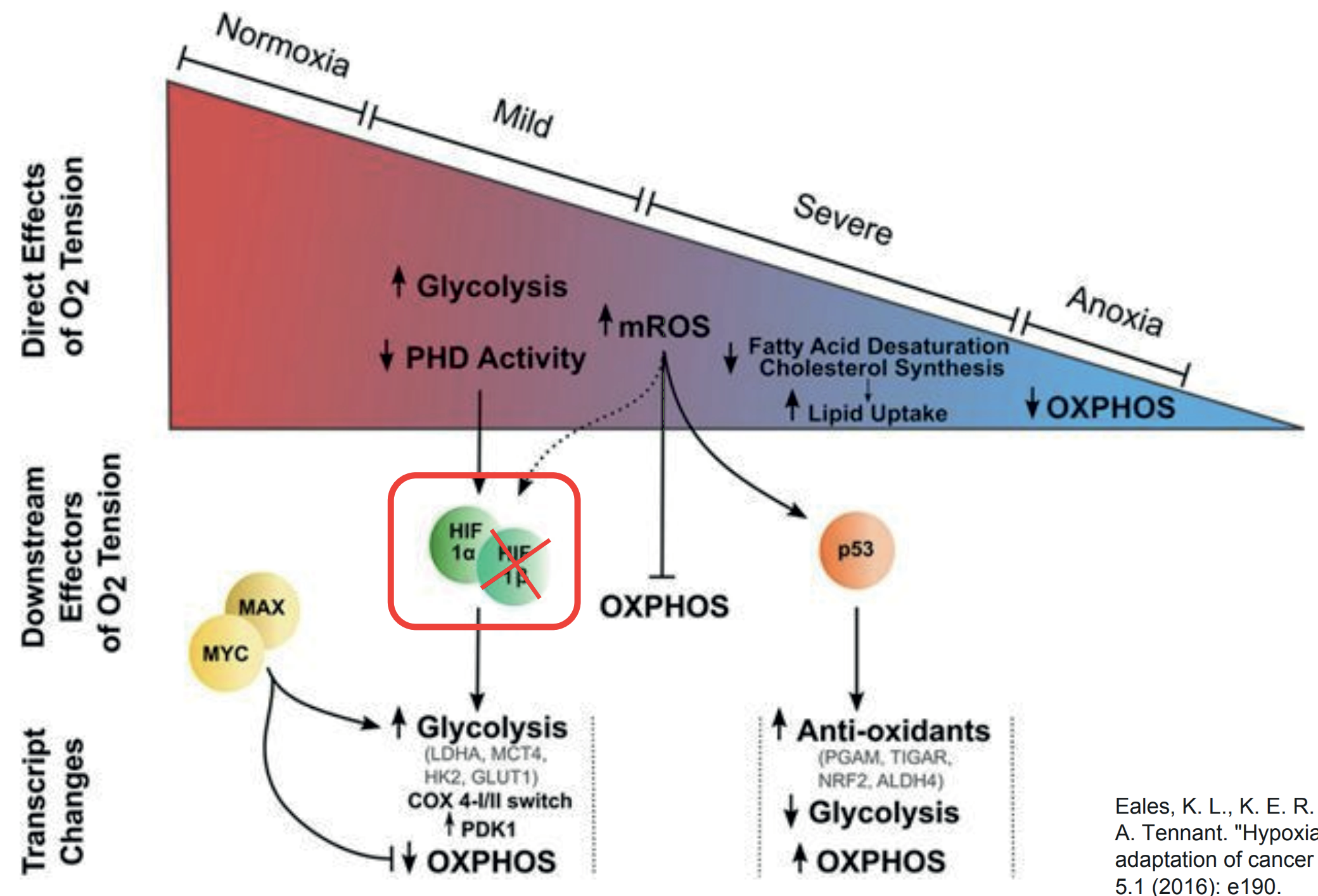
Identification and Characterization of HIF-Dependent Altenative Splicing Events in Pancreatic Cancer
( link to paper )
